NGS · Snakemake · Python
Sequana Project
a set of NGS pipelines (sequence analysis)
ATCGATGCTAGCTAGCATGCTA
Sequana: a set of NGS pipelines
This page serves as an additional set of resources for
the Sequana project. Sequana is comprised of a Python library, the
documentation of which is accessible on Read The Docs.
The source code for the Python library can be found on
GitHub.
If you use Sequana, please cite: Cokelaer et al.,
Journal of Open Source Software, 2017
(doi:10.21105/joss.00352).

One of the primary objective of the Sequana project is to offer a collection of NGS pipelines.
The pipelines currently available are located on PyPI and the Sequana GitHub organisation page,
each having their own repository. For instance, the RNA-seq pipeline can be found at sequana/rnaseq.
This page is also a place for providing supplementary information to the main repositories.
Pipelines
Sequana provides a collection of NGS pipelines. Each pipeline lives in its own repository on the
Sequana GitHub organisation. Below are some highlighted pipelines:
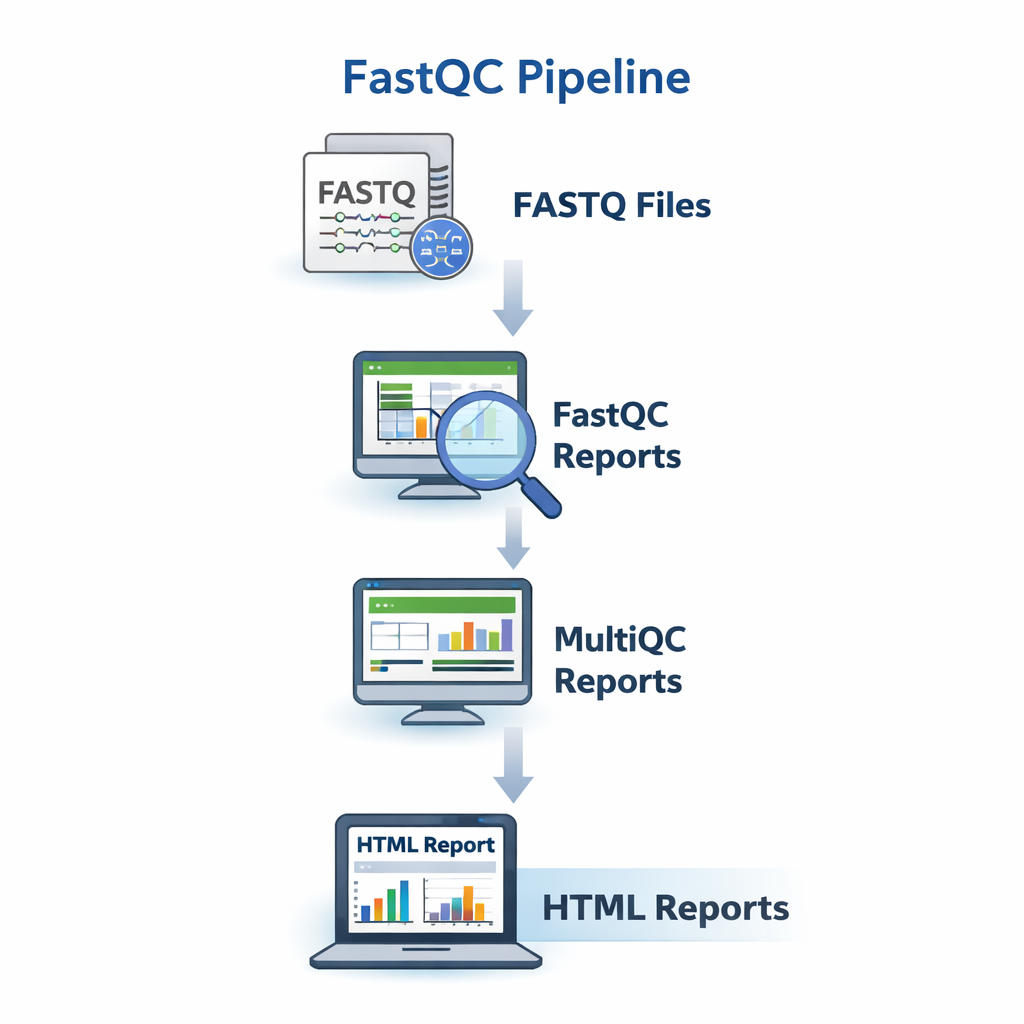
sequana_fastqc
A quality-control pipeline that runs FastQC and MultiQC on raw FASTQ files to assess sequencing quality.
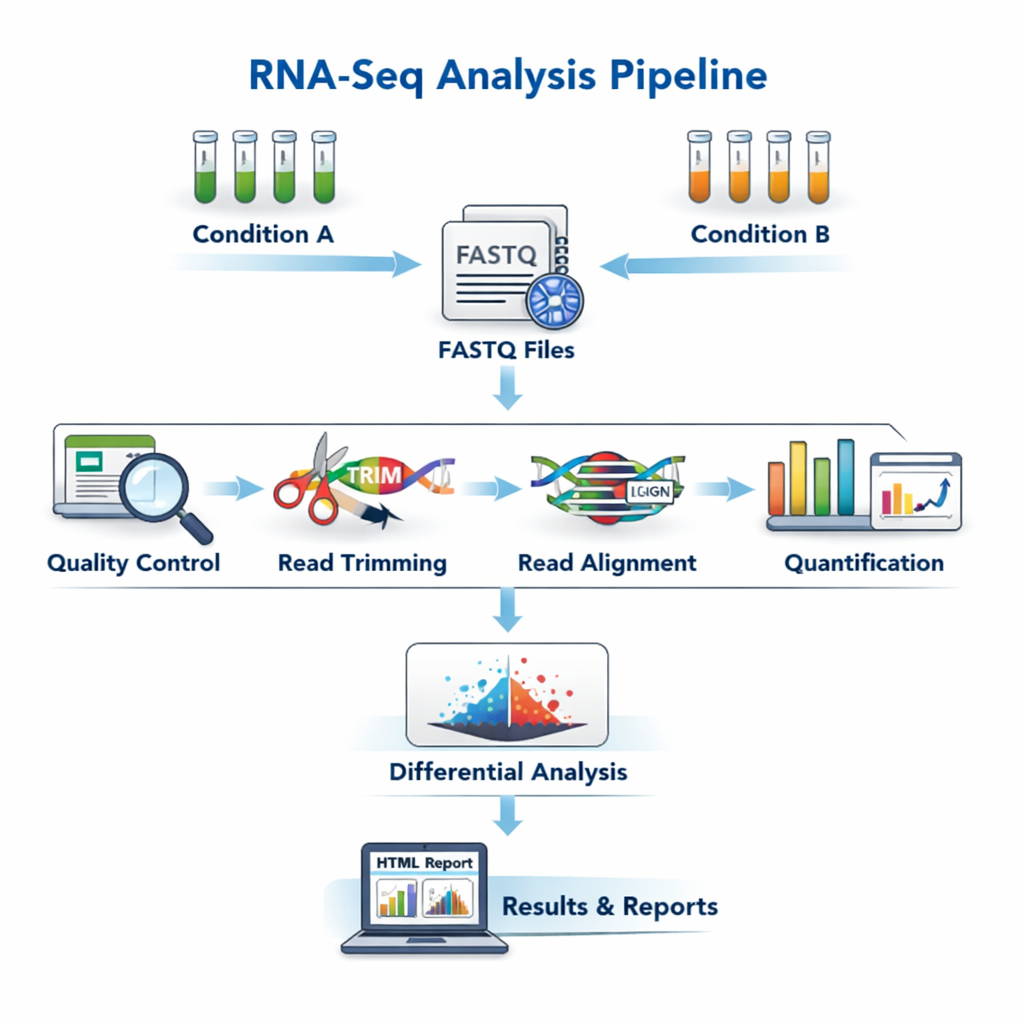
sequana_rnaseq
An RNA-seq pipeline that covers trimming, alignment, feature counting, and differential expression analysis.
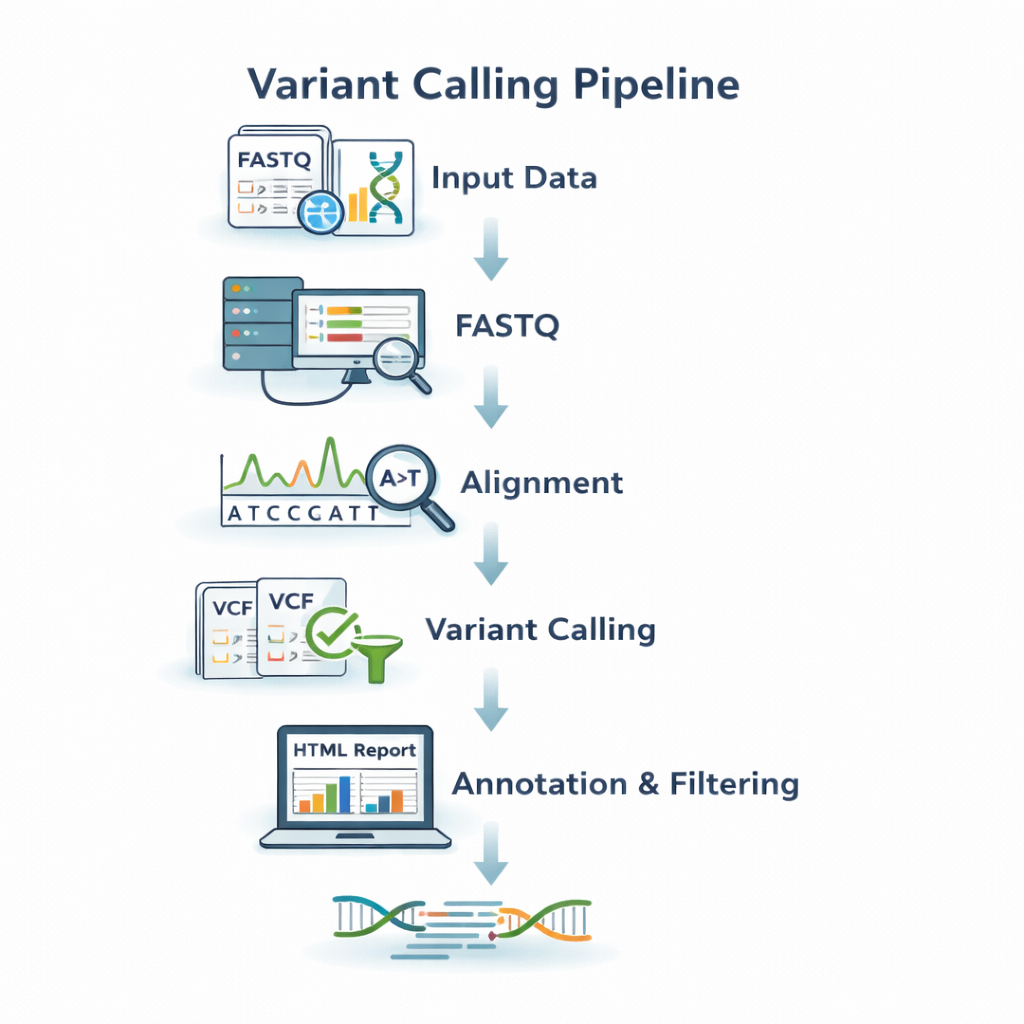
sequana_variant_calling
A variant-calling pipeline that performs read trimming, alignment, and SNP/indel detection
using standard tools such as BWA and freebayes, with interactive HTML reports.
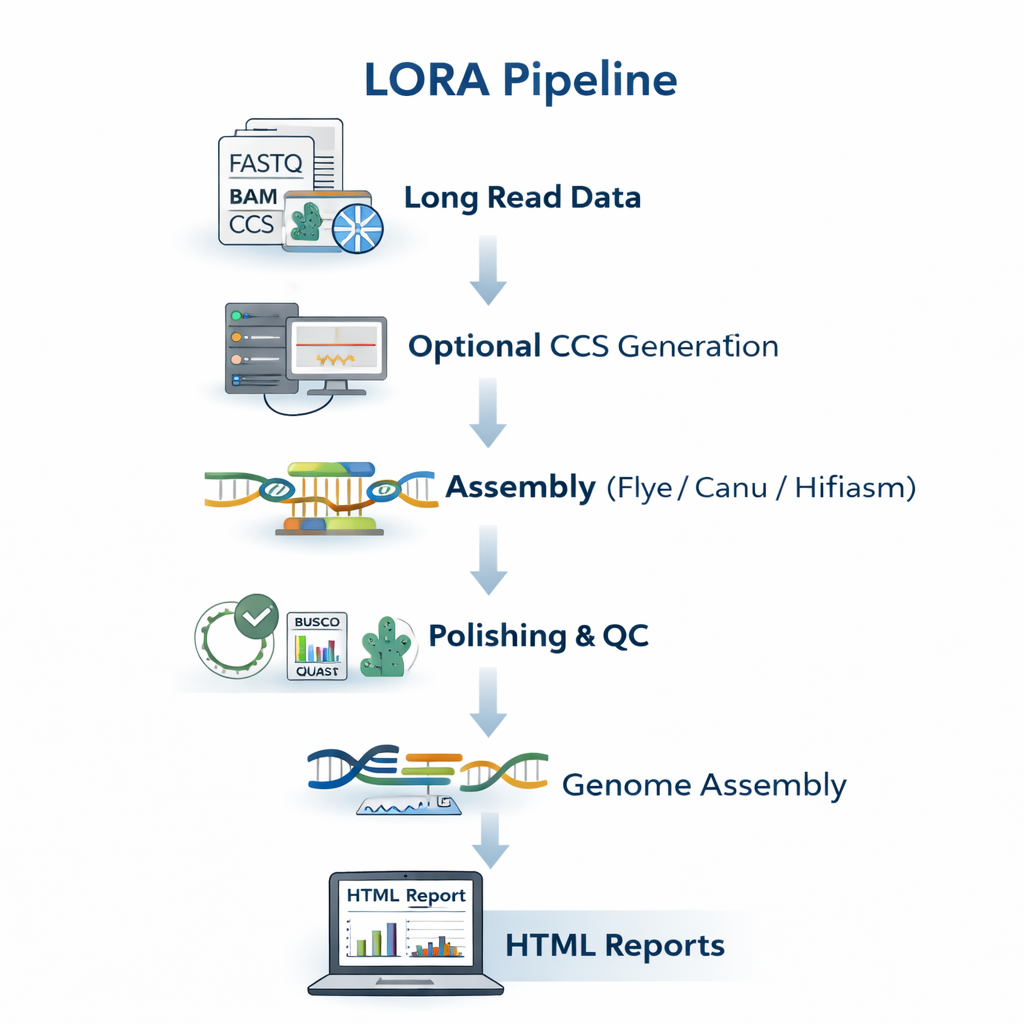

sequana_lora
A long-read assembly pipeline for Oxford Nanopore (and PacBio) data that performs
basecalling, quality control, de-novo assembly, polishing, and assembly quality
assessment with tools such as Flye, Medaka, and BUSCO.
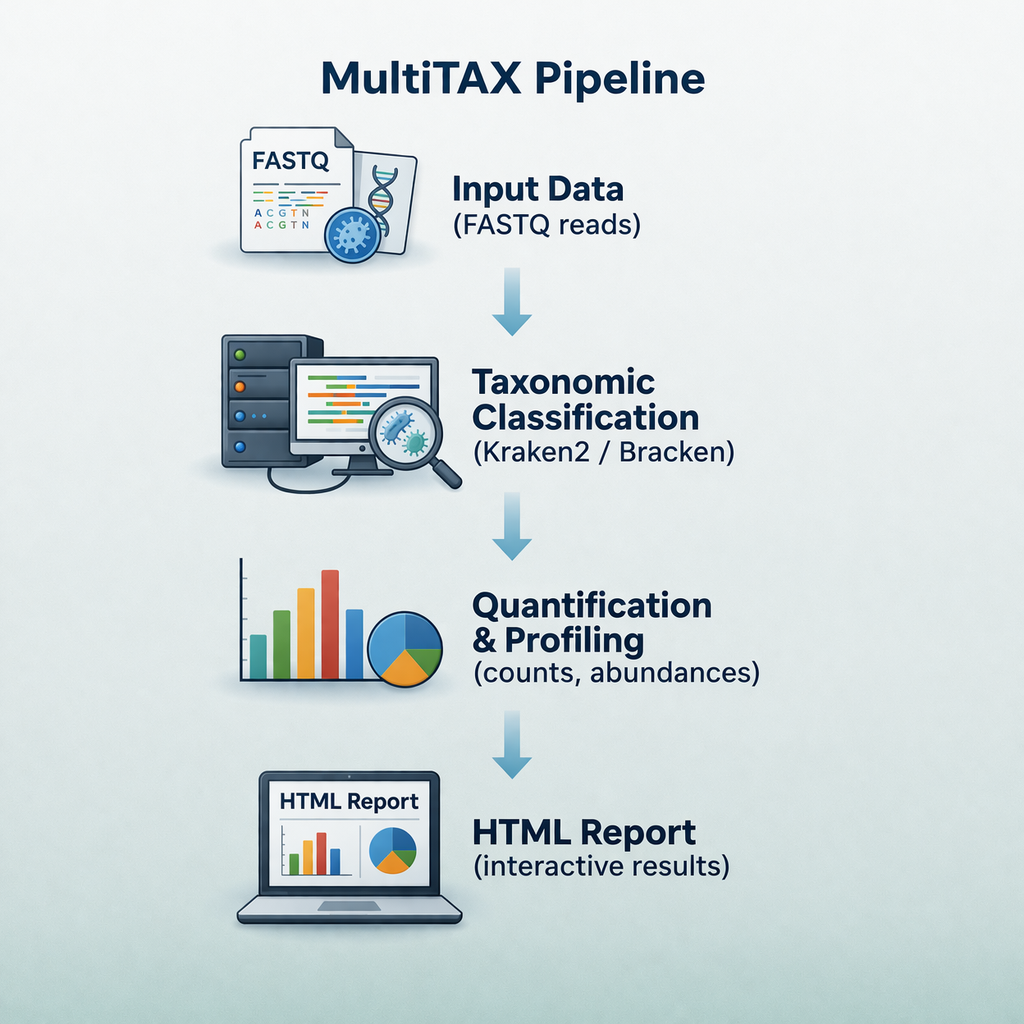
sequana_multitax
A taxonomic classification pipeline that runs multiple classifiers (Kraken2, Centrifuge, etc.)
on sequencing reads, merges results, and produces comparative summary reports.
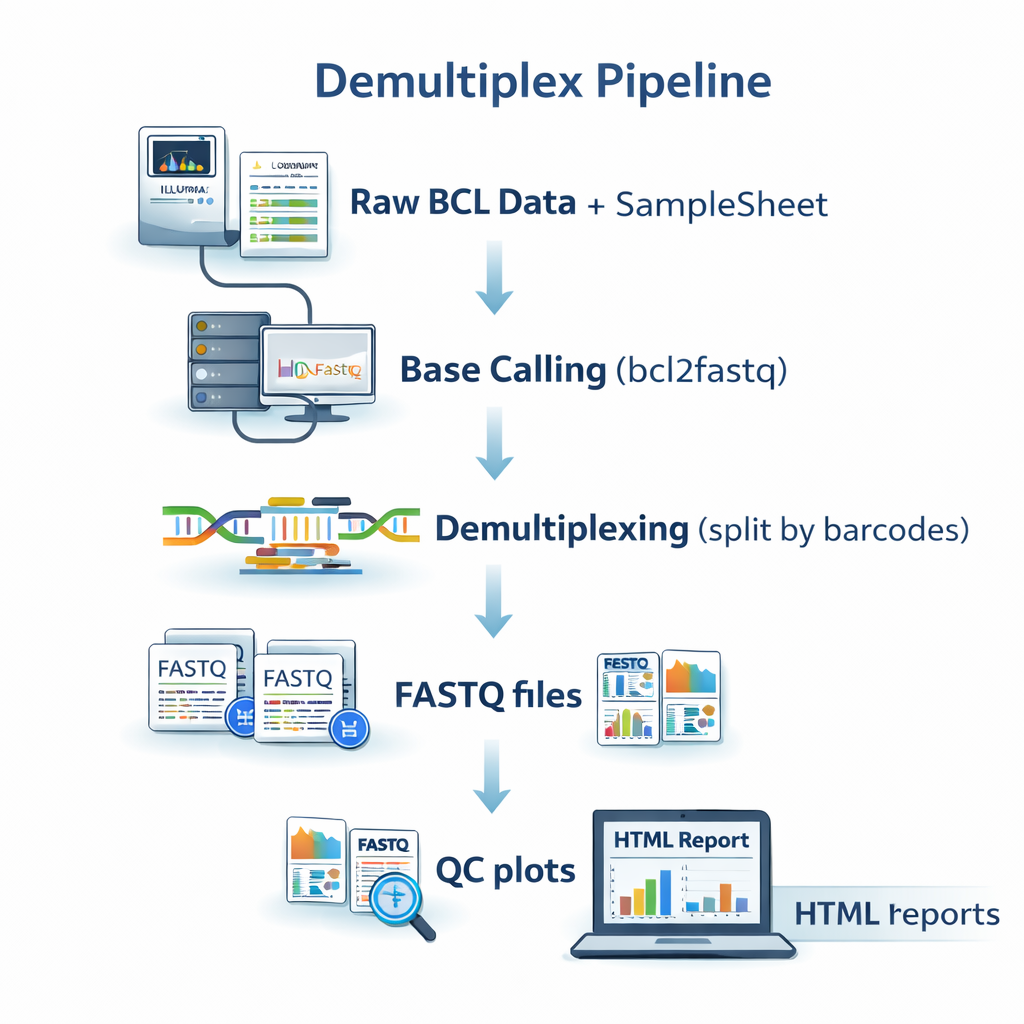
sequana_demultiplex
A demultiplexing pipeline built around bcl2fastq / bcl-convert that converts Illumina
BCL files into FASTQ files, validates sample sheets, and generates MultiQC summary reports.
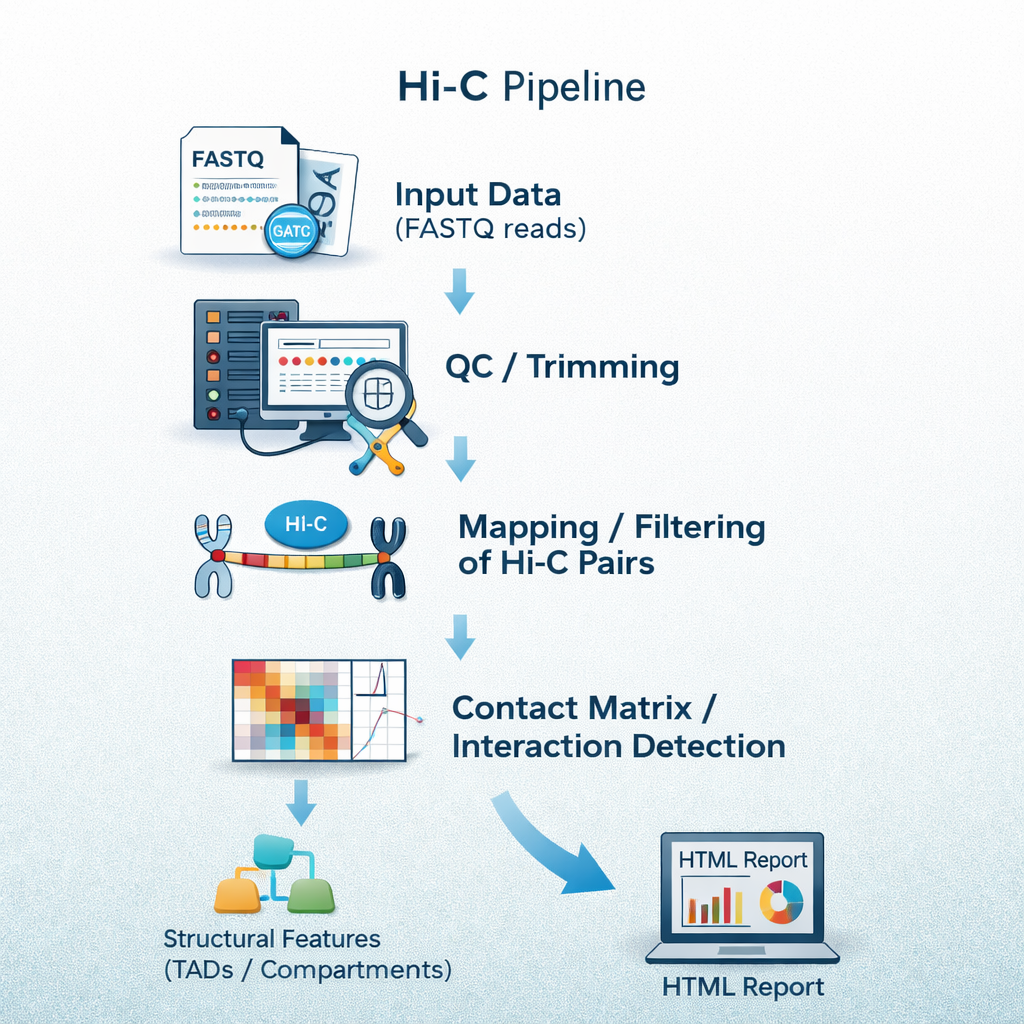
sequana_hic
A Hi-C pipeline for chromatin conformation capture data that performs read trimming,
alignment, contact-map generation, and quality assessment of 3D genome organisation.

sequana_chipseq
A ChIP-seq pipeline that covers read trimming, alignment, peak calling, and
downstream annotation to identify protein–DNA binding sites and histone modifications.
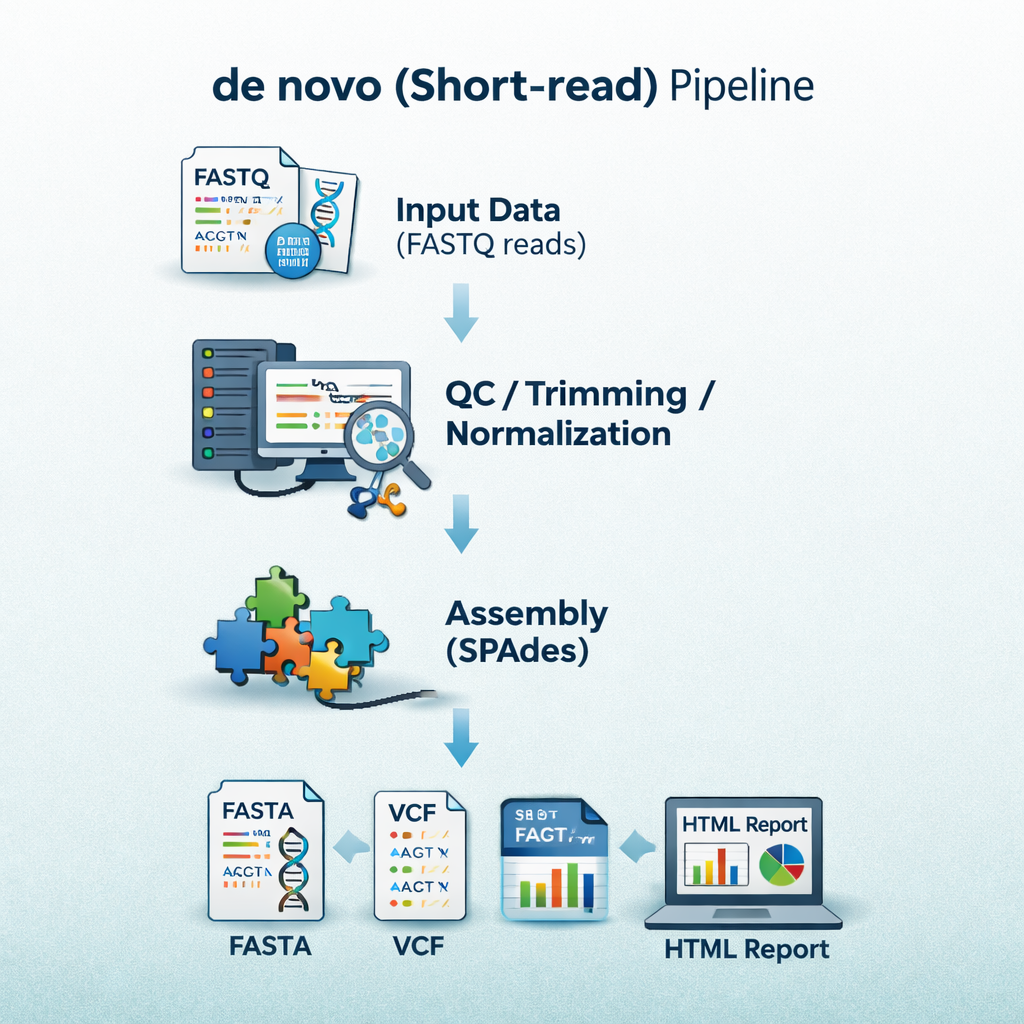
sequana_denovo
A de-novo assembly pipeline for short-read Illumina data that performs trimming,
assembly, scaffolding, and assembly quality assessment with tools such as SPAdes and QUAST.
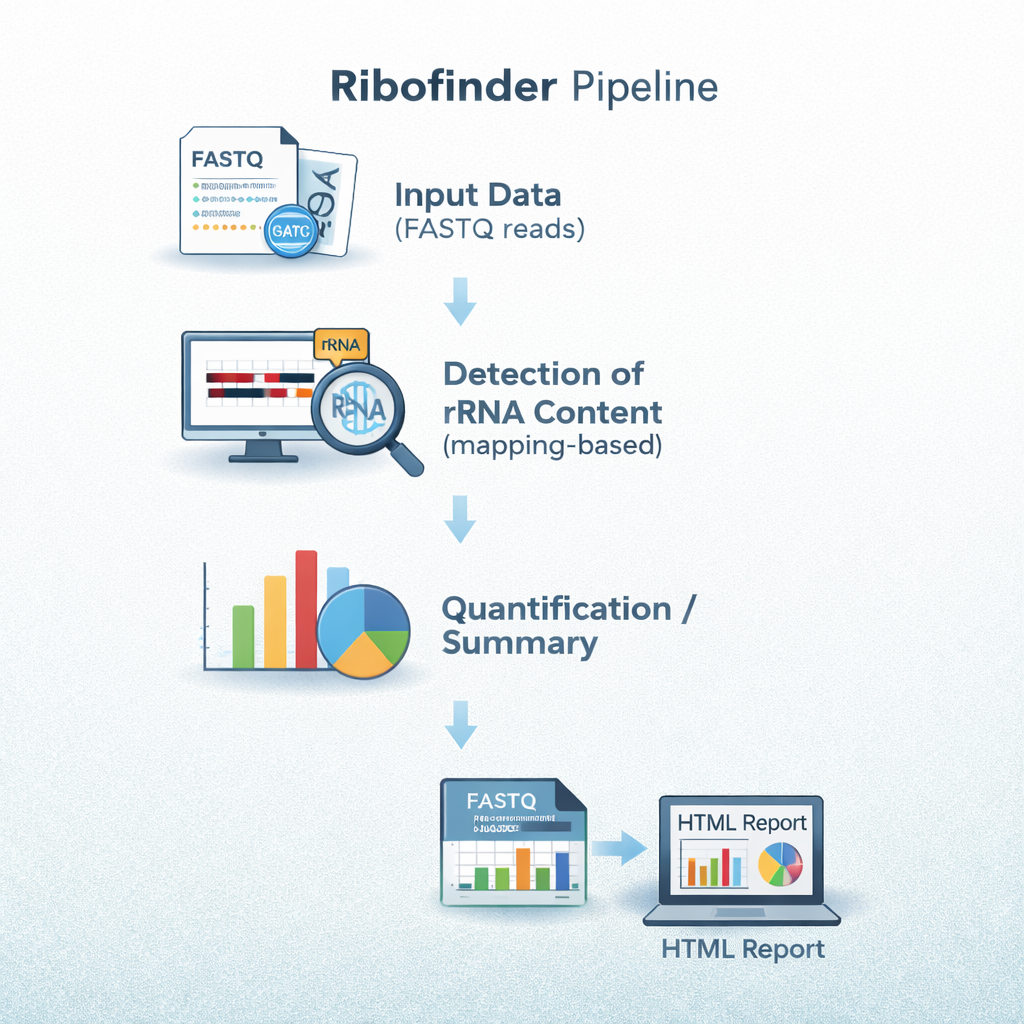

sequana_ribofinder
A pipeline that quantifies ribosomal RNA content in sequencing data by mapping reads
against rRNA references, helping to assess rRNA contamination and depletion efficiency
in RNA-seq experiments.
These are just a few highlights. Sequana encompasses many more pipelines — both public and
private — covering a wide range of sequencing applications. A broader overview is available on
github.com/sequana/sequana,
the Sequana GitHub organisation,
and the full documentation at
sequana.readthedocs.io.
Sequana ships published tools and a set of standalone sub-commands invocable directly from
the command line (sequana <subcommand>), all distributed as part of the
sequana Python package.
Talks
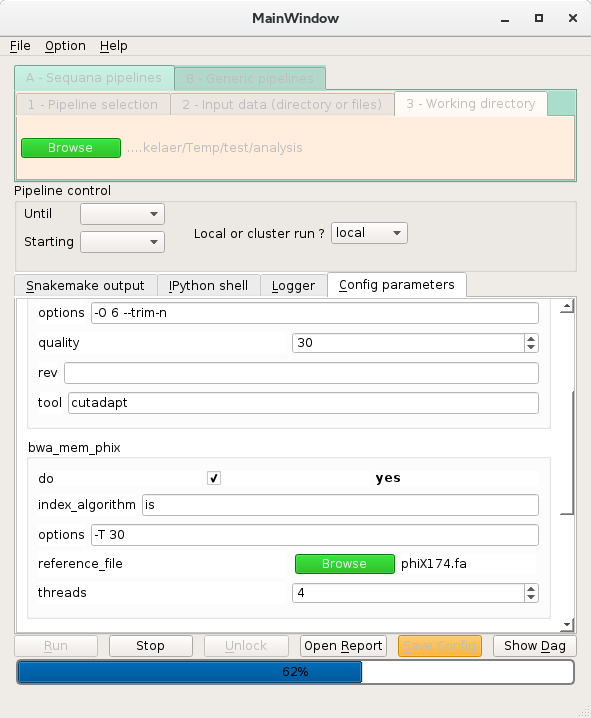
Web Applications
Check your Illumina Sample Sheet
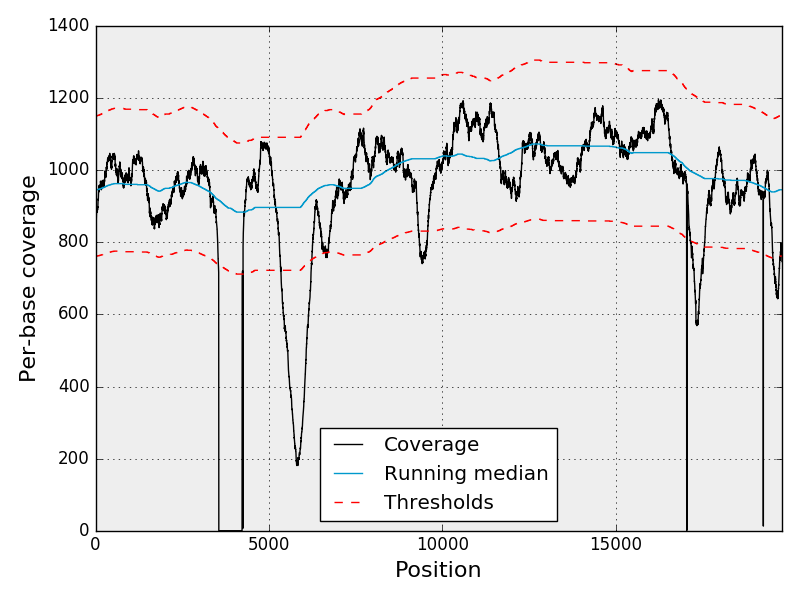
Report examples (click on the image)
The following was generated with the Sequana Coverage standalone application and keep here a it is reference in other places. This is the coverage along a virus genome where the black dot line represent the coverage, the blue is the median coverage, and red lines are the top and bottow thresholds for detecting event of interest.